From a very basic perspective, DNA sequencing definition is determining the order of the nucleotides in a person’s DNA. To be more specific, it involves reading off the As, Ts, Cs, and Gs that make up the sequence of a person’s genome.
What is DNA sequencing?
When put together, these four letters spell out an individual’s complete genetic code.
In other words, DNA sequencing tells us the unique story that is written in each person’s genes. And because every individual has a slightly different genome sequence,DNA sequencing can be used to distinguish one person from another.
It can also help us understand why people look and act differently from one another—or why they might be susceptible to certain diseases.
How does DNA sequencing work?
By looking at the DNA sequence of people with a particular disease, scientists can often identify genetic changes that might be responsible for triggering the disorder.
In recent years, DNA sequencing has become much more widespread as the technology necessary to do it has become cheaper and more readily available. This has revolutionized medicine and biology, as researchers can now quickly and easily sequence genomes from all sorts of organisms, including humans.
DNA sequencing is not to be confused with genetic testing, which looks for specific changes in an individual’s DNA that are known to be associated with disease. Most DNA sequencing tests do not look for specific diseases but rather provide a person with information about their complete genome.
Steps of DNA Sequencing
DNA Sequencing is the process of determining the precise order of nucleotides within a DNA molecule. It’s a key tool in genetics, because it allows scientists to identify changes in genes that might be associated with specific traits or health conditions. Sequencing can also be used to create so-called “DNA fingerprints,” which can be used forensically to identify individuals.
What is sequencing and how is it used?
There are several different methods for sequencing DNA, but they all essentially involve breaking up the double strands of DNA, reading the order of bases on one strand, and then using computers to piece together that information into a longer sequence.
Here we’ll focus on two popular sequencing methodologies: Sanger sequencing and next-generation sequencing (NGS).
Sanger Sequencing
The Sanger method, also called chain termination sequencing, was developed in 1977 by Frederick Sanger and colleagues at Cambridge University.1 It quickly became the standard methodology for DNA sequencing and remained so for nearly 30 years. The basic principles are still used today in what’s known as dideoxy sequencing (or ddRADseq). In this approach, DNA fragments are generated by randomly shearing genomic DNA (this can be done chemically or with physical methods like sonication).
The fragments are then ligated to each other and inserted into a sequencing vector, which is typically a plasmid. Next, the DNA construct is introduced into bacteria, where it replicates. Because the DNA template has random shearing sites, the insert will have different lengths in different bacterial colonies.
To sequence the insert, special primers are designed that will anneal (or bind) to specific regions flanking the insert. For example, if the plasmid used for cloning has an M13 origin of replication, primers that anneal to this region can be used. PCR is then used to amplify the region between the two primers (i.e., the insert), using dNTPs that also contain one of four modified bases—A, C, G, or T—at their 3′-end known as chain terminators.
When these modified nucleotides are incorporated into newly synthesized DNA strands during PCR amplification, they terminate strand extension because their 3′-OH group has been replaced by another group that cannot form a phosphodiester bond with the next nucleotide in the template strand.
As a result, PCR amplification results in a mixture of DNA fragments of different lengths—each terminated by one of the four chain terminators.
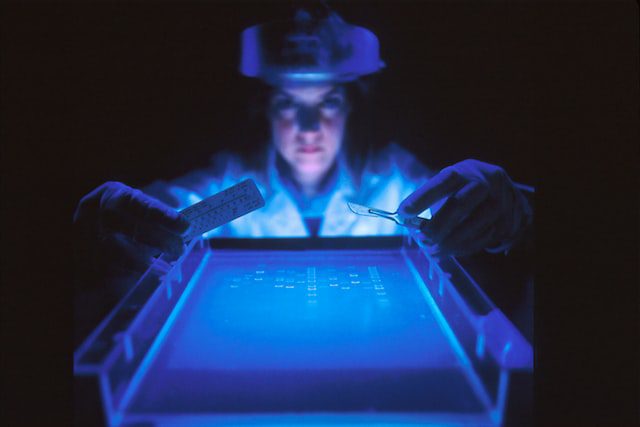
The mix of DNA fragment sizes is then separated by electrophoresis on an agarose gel, and the resulting “ladder” is visualized by staining with ethidium bromide (see image below). The position of each band corresponds to a particular DNA fragment length. Finally, the ladder is sequenced using the original primer pairs, and the order of bases is read from bottom to top.
Next-Generation Sequencing (NGS)
The Sanger sequencing method was relatively slow and expensive, so scientists began looking for ways to speed up the process and reduce costs. This eventually led to what’s now called next-generation sequencing (NGS), which can generate millions or even billions of sequences in a single run. As a result, NGS has become the preferred approach for many applications, including genome sequencing projects like those conducted by The Cancer Genome Atlas (TCGA) and The Human Genome Project.
Like Sanger sequencing, NGS also involves randomly shearing genomic DNA and using PCR to amplify fragments for sequencing. However, there are several key differences in the methodology. Perhaps the most important is that NGS uses much shorter read lengths—typically around 100-300 bases. This is possible because NGS machines can rapidly generate large numbers of short sequences (called reads) in parallel.
In addition, NGS machines don’t use chain terminators like the Sanger method; instead, they incorporate fluorescently labeled nucleotides at specific positions along the template DNA strand. As each position is sequenced, a particular color of light is emitted. By reading these colors in order, it’s possible to determine the sequence of the original template strand.
NGS machines can generate millions or even billions of sequences in a single run, making them much faster than Sanger sequencing machines.11 In addition, because NGS doesn’t require costly chain terminators or special primers, it’s less expensive than Sanger sequencing—making it more accessible to many laboratories and researchers.
However, NGS also has some disadvantages. Because reads are shorter, they’re more likely to contain errors.
In addition, NGS machines are more complex and expensive than Sanger sequencing machines.14 As a result, Sanger sequencing is still used for some applications—particularly when longer read lengths or higher accuracy is required.
DNA sequencing alignment
In order to sequence a DNA molecule, it is necessary to first remove it from the chromosome and cut it into smaller pieces. Each of these smaller pieces is then inserted into a sequencing machine, which will read the order of the nucleotides in that particular piece of DNA.
In order to obtain an accurate sequence, it is often necessary to align the different pieces of DNA that have been sequenced. This process is known as DNA sequencing alignment.
There are numerous methods that can be used for DNA sequencing alignment, including:
Local alignment: This method compares short segments of two sequences in order to find regions of similarity between them. This is typically done using a software program such as BLAST or FASTA.
Global alignment: This method compares the entire length of two sequences in order to find the best overall match between them. This is typically done using a software program such as Needleman-Wunsch or Smith-Waterman.
Multiple sequence alignment: This method compares multiple sequences at once in order to find similarities between them. This is typically done using a software program such as CLUSTAL or MAFFT.
DNA sequencing alignment is crucial tool for biologists as it can be used to help understand the evolutionary relationships between different species. It can also be used to identify functional regions of DNA, or to find mutations that may be responsible for disease.
Future of DNA sequencing
The future of DNA sequencing is immensely exciting. It’s now possible to sequence an entire human genome for less than $1,000, and the speed and accuracy of sequencing continues to improve. This technology is being used in a variety of ways, from identifying genetic diseases to finding new cancer treatments.
DNA sequencing has already had a huge impact on medicine. We can now diagnose genetic diseases with much greater accuracy and efficiency than ever before. And as our understanding of the genome improves, we will be able to develop targeted therapies for a wide range of conditions.
Cancer research is another area where DNA sequencing is having a major impact. By understanding the unique mutations that drive each person’s cancer, we can develop personalized treatments that are much more effective than traditional chemotherapy. We are also using DNA sequencing to screen for potential cancer-causing genes, so that we can take preventive measures against this disease.
The field of forensics is also benefiting from DNA sequencing technology. Crime scene investigators can now quickly identify unknown individuals by their DNA profile, making it possible to solve crimes that would otherwise remain unsolved. In the future, DNA profiling may even be used for routine security screening, such as at airports or government buildings.
As DNA sequencing technology continues to improve, its applications will become even more widespread. We will have a better understanding of human evolution and the genetic basis of disease. We will be able to create personalized medication and tailor cancer treatments to each individual patient. And we will be able to solve crimes that would otherwise remain unsolved. The future of DNA sequencing is immensely exciting, and the possibilities are endless.